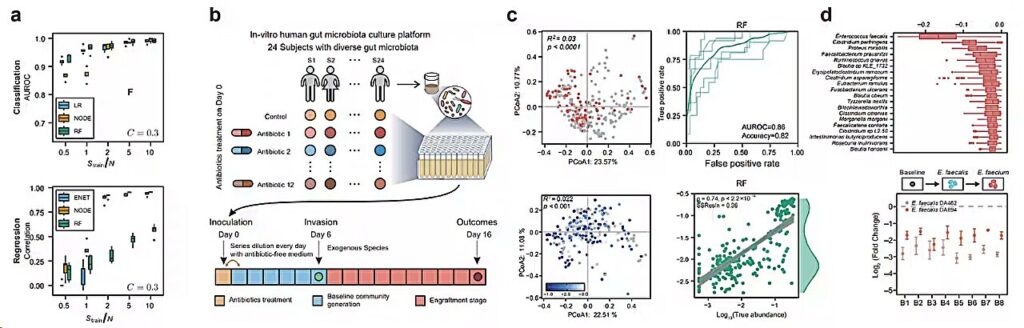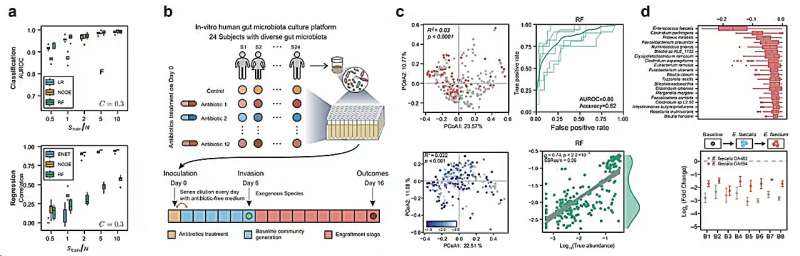
Data-driven prediction of colonization outcomes for complex microbial communities. Credit: Dai Lei.
Microbial communities are constantly exposed to invasion by alien species, which can significantly alter their composition and function. The ability of microbial communities to resist invasion is thought to be an emerging property that arises from complex interactions between component species.
The ability to predict and modify colonization outcomes (i.e. to prevent pathogen implantation and promote probiotic implantation) is crucial for microbiota-based personalized interventions in nutrition and medicine. Despite accumulating empirical studies, predicting colonization outcomes in complex communities remains a fundamental challenge due to limited knowledge about interspecies interactions.
Recently, a research team led by Professor Dai Lei from the Shenzhen Institute of Advanced Technology (SIAT), Chinese Academy of Sciences, in collaboration with other researchers, developed a dynamic model-independent, data-driven approach to predict the colonization outcomes of invasive species in complex microbial communities, even without detailed knowledge of the underlying ecological and biochemical processes.
This study Nature Communications March 16th.
In this study, the researchers systematically evaluated the proposed data-driven approach using classical ecological dynamic models and synthetic data generated from in vitro human fecal-derived microbial communities. They found that with sufficient sample sizes of training data, [on the order of ~O(N)]Machine learning models can be used to predict colonization outcomes (i.e. whether an invasive species is able to establish and, if so, its abundance).
The researchers then generated a large dataset containing in vitro experimental results for two representative species that colonize microbial communities derived from human feces. They also validated that their machine learning model could predict experimental colonization outcomes (AUROC > 0.8).
Additionally, the researchers used machine learning models to identify species that have a significant impact on colonization, demonstrating that the introduction of highly interacting species can significantly alter colonization outcomes.
“Our findings demonstrate that the colonization outcomes of complex microbial communities can be predicted and tuned through data-driven approaches,” Professor Dai said.
“Data-driven methodologies are powerful tools for biologists. Combined with advances in predicting the properties of complex biomolecules, we expect this approach will bring a paradigm shift in the study of the stability and function of complex ecosystems and facilitate important applications in medicine and agriculture.”
More information:
Lu Wu et al., Data-driven prediction of colonization outcomes of complex microbial communities, Nature Communications (2024). DOI: 10.1038/s41467-024-46766-y
Provided by Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences
Citation: Machine learning models can predict colonization outcomes of complex microbial communities (August 29, 2024) Retrieved August 29, 2024 from https://phys.org/news/2024-08-machine-colonization-outcomes-complex-microbial.html
This document is subject to copyright. It may not be reproduced without written permission, except for fair dealing for the purposes of personal study or research. The content is provided for informational purposes only.