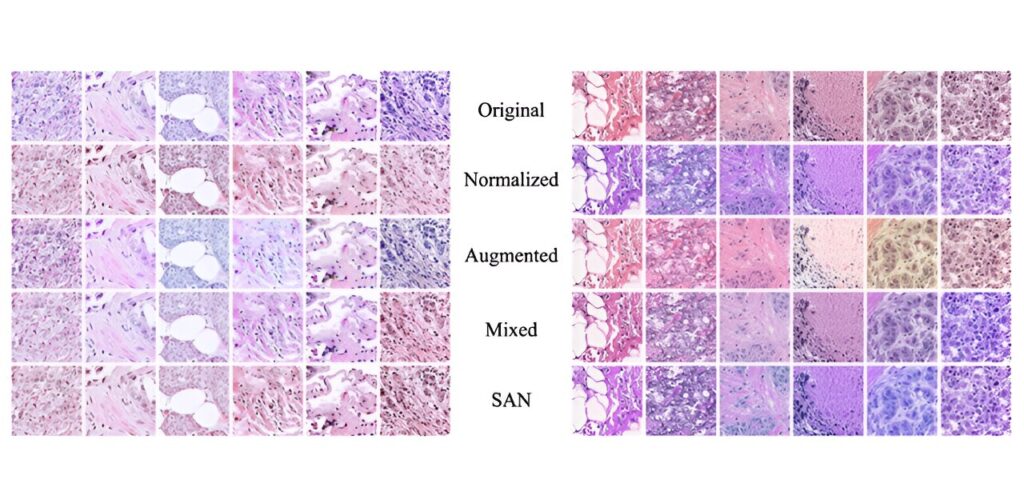
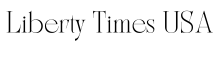The stained SANS processed tissue image dataset (bottom row) achieves a more consistent color distribution than those processed with other techniques. Such consistency is essential when training machine learning based systems. Credit: Medical Imaging Journal (2024). DOI: 10.1117/1.JMI.11.4.044006
In histopathology, where tissues are studied under a microscope to understand and diagnose disease, stains are essential tools. Simply put, stains are carefully selected or created chemicals that attach to specific cellular components. When viewed under a microscope, they change the observed color, helping users to more easily identify cellular structures.
Datasets of stained images showing both normal and diseased tissues are useful for training machine learning models, helping doctors evaluate difficult cases and reducing personal biases when diagnosing. To ensure these models perform well, it is crucial to minimize the color differences between the images used for training and those analyzed in real-world scenarios. So-called “domain adaptation techniques” can be used to correct color variations resulting from the unique experimental setups used in different laboratories, resulting in more consistent and comparable data.
In a recently published study, Medical Imaging JournalResearchers from the University of North Carolina at Chapel Hill (USA) have proposed a new domain adaptation technique. Called Stain SAN, the proposed method could help make stained histopathological image datasets more useful to many emerging machine learning-based classification systems, ultimately leading to improved diagnostic tools.
Stain SAN has three main steps: stain extraction, color adaptation, and intensity adaptation. In the first step, the original stained image is decomposed into the product of two matrices: one containing color information and the other containing light intensity information for each pixel. In the adaptation step, the distribution of colors in the color matrix is modified through a statistical process that takes into account the training images, ensuring that the modified colors fall within the target distribution. Finally, in the third step, before image reconstruction, the intensity matrix is randomly varied, which increases the diversity of the stain domain.
“The main advantage of Stain SAN is that it combines the strengths of traditional staining adaptation methods while overcoming their inherent weaknesses,” explains lead researcher Taebin Kim, PhD. “Other established techniques, such as staining standardization, staining enhancement, and staining mixing, can be understood as special cases of Stain SAN.”
The researchers tested their approach qualitatively and quantitatively using histopathological images from publicly available datasets. Based on their observations and feedback from expert pathologists, the researchers found that stained areas were more generalized and colors aligned more consistently in image datasets processed with Stain SAN. Additionally, Stain SAN enhanced the contrast between the nucleus and cytoplasm of each cell, highlighting the differences between tumor cells and supporting tissue.
The researchers also trained a machine learning-based classifier using the dataset processed with various domain adaptation techniques and tested its performance on images processed from a different dataset. Interestingly, Stain SAN outperformed all the aforementioned methods, achieving significantly higher accuracy.
“Our results clearly demonstrate the improvements made throughout the history of the development of these methods, culminating in the significant enhancements brought by Stain SAN,” commented Kim. “Furthermore, we showed that Stain SAN can achieve results comparable to state-of-the-art deep learning-based approaches without requiring separate training for stain adaptation or access to the target domain during training, which is unrealistic in clinical settings. This highlights the effectiveness and computational efficiency of Stain SAN.”
The development of efficient domain adaptation techniques such as Stain SAN is essential to bridge the gap that exists between machine learning systems and their applications in medicine. The research team is already planning potential improvements to the method and conducting further tests with other datasets.
“Our findings confirm that Stain SAN is a robust approach for stain region adaptation in histopathology images, leading to advances in this field of computational tasks,” Kim concludes, expressing optimism for the future.
These efforts will result in more accurate and convenient diagnostic protocols, saving time for both doctors and patients.
More information:
Taebin Kim et al. “Stain SAN: Simultaneous Enhancement and Normalization of Histopathology Images” Medical Imaging Journal (2024). DOI: 10.1117/1.JMI.11.4.044006
Citation: Color-adjusting technique for histopathology image datasets could help enhance machine learning-based diagnostic tools (August 26, 2024) Retrieved August 26, 2024 from https://medicalxpress.com/news/2024-08-adjusting-technique-histopathology-image-datasets.html
This document is subject to copyright. It may not be reproduced without written permission, except for fair dealing for the purposes of personal study or research. The content is provided for informational purposes only.