patient
The Institutional Review Board of the Chronic Corona Treatment Center reviewed and approved the protocol. All participants provided written informed consent to participate in the study. The date of acute corona infection was defined as the date of the first positive SARS-CoV-2 test result or the onset of corona symptoms. Healthy control participants had no history of SARS-CoV-2 infection at the time of enrollment and had negative anti-NP tests. All methods were performed in accordance with the relevant guidelines and regulations set forth by the review board. All patients were aged 18 years or older.
Mild acute COVID-19:
-
1.
Fever, cough, sore throat, fatigue, headache, muscle pain, nausea, diarrhea, loss of taste or smell,
-
2.
Chest imaging (CXR or chest CT) shows no signs of pneumonia;
-
3.
There is no shortness of breath or difficulty breathing.
Moderate acute COVID-19:
-
1.
radiological evidence of pneumonia, fever and respiratory symptoms,
-
2.
Oxygen saturation (SpO2) in indoor air at sea level is 94% or greater.
LC/PASC
Inclusion criteria for individuals in the LC group were previous confirmed or suspected COVID-19 infection (per World Health Organization guidelines), age ≥18 years, and symptoms persisting for ≥12 weeks after initial COVID-19 infection. Symptoms included those previously described and scored.4.
Inclusion criteria for healthy controls (HCs) were age ≥18 years, no previous SARS-CoV-2 infection, and a negative medical history performed as part of enrollment in a Chronic COVID Treatment Center (CCTC).
Chronic Lyme Disease (CLD)
The patient presented to the CCTC with a history of fatigue, brain fog, and post-exertional malaise lasting for more than 6 months since before the SARS-CoV-2 pandemic (pre-2020) (per the ILADS Working Group).13.
The existence of Borrelia burgdorferi Confirmation was achieved by a two-step immunological test, including immunoblotting. The presence of other tick-borne organisms was noted but, as previously mentioned, was not required for the definition of CLD.13.
Multiplex cytokine/chemokine profiling
Plasma collected in plasma preparation tubes (PPT, BD Biosciences, San Jose, CA) was used for cytokine quantification using a customized 14-plex bead-based flow cytometry assay (IncellKINE, IncellDx, Inc) on a CytoFlex flow cytometer, as previously described, with the following analytes: TNF-a, IL-4, IL-13, IL-2, GM-CSF, sCD40L, CCL5 (RANTES), CCL3 (MIP-1a), IL-6, IL-10, IFN-g, VEGF, IL-8, and CCL4 (MIP-1b).2For each patient sample, 25 μL of plasma was used for each well of a 96-well plate. Samples were analyzed on a Beckman Coulter CytoFlex LX 3-laser flow cytometer using Kaluza Analysis Software (Beckman-Coulter, Miami, FL). All statistical analyses were performed using the Mann–Whitney test, and a P value ≤ 0.05 was considered statistically significant.
Data acquisition and processing for building machine learning models
To construct the working dataset, we selected cytokine profiles from three disease states: non-impaired (NP), LC/PASC, and CLD. The non-impaired class represents the collection of unaffected (healthy controls) and mild-to-moderate COVID-19 patients. We combined categories based on the absence of chronic immunological impairment, as previously published.4 It is also present in chronic disease states such as PASC and CLD. When comparing IncellKine cytokine profiles, there is no statistical difference between the two states (pThe relative values of the mutagenic proteins using the Mann-Whitney U test (-value > 0.05) also support this classification. Severe COVID-19 patients, i.e., COVID-19 infected individuals presenting severe symptoms and immunological fluctuations, were excluded. Outliers were removed using isolation forest (contamination parameter = 5%), generating a dataset consisting of 67 no fluctuations, 103 PASC, and 53 CLD. Each individual had a cytokine profile obtained from the incellKINE assay (14-plex cytokine panel), LHI (Long-Term Infected Index), and SI (Severity Index) calculated according to equations (1) and (2) as reported in the literature.2:
$$LHI= \frac{IL-2+IFN-gamma}{CCL4}$$
(1)
$$SI= \frac{ \left(IL-6+\frac{sCD40L}{1000}+\frac{VEGF}{10}+(10*IL-10\right)}{\left(IL-2 + IL-8\Right)}$$
(2)
The dataset was then imported into Python using the Pandas library.14,15,16Data were stratified using the train_test_split function in the model_selection module sci-kit-learn.1780% of the data was for training, and a 20% holdout evaluation split was used to obtain performance metrics and identify overfitting. Table 1 contains the number of instances in the dataset, training, and evaluation partitions before splitting.
Building tree-based machine learning classifiers: decision trees, random forests, gradient boosting machines
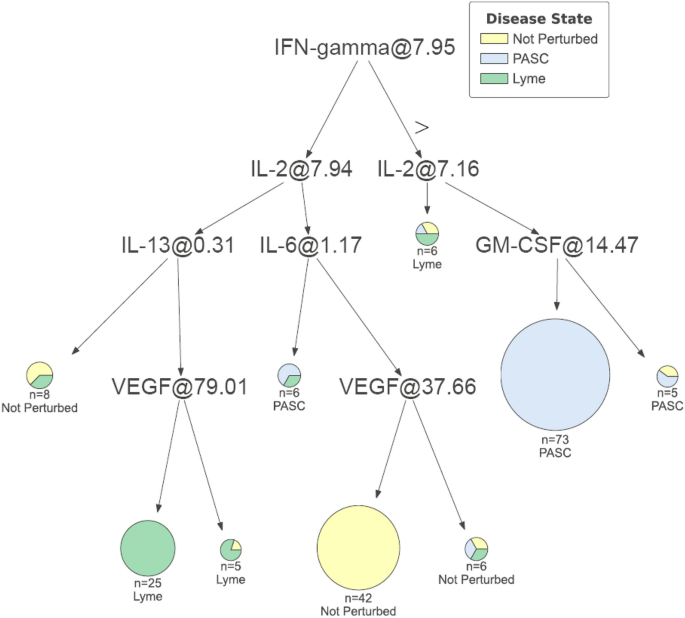
In this study, we employed three tree-based machine learning classifiers: decision tree, random forest, and gradient boosting machine. Decision tree and random forest were implemented using the sci-kit-learn library, while gradient boosting machine utilized the LightGBM library. Optimizing the hyperparameters of each model involved different settings. For decision tree, parameters such as criterion, class weights, splitter, max depth, min sample split, and leaf were tuned. For random forest model parameters included number of estimators, criterion, max depth, min sample split and leaf, and bootstrap options. For gradient boosting machine, we modified learning rate, number of estimators, min data for leaf, and depth. Hyperparameter tuning was performed with three iterations of 10-fold cross-validation and the best model was selected based on F1 score. Performance was evaluated with a 20% holdout evaluation split. We calculated performance metrics using a custom classification report including recall, specificity, precision, negative predictive value, and F1 score to determine if there was overfitting of the model. The best performing model was assigned to the best_model variable.
Development of a Lyme disease index to further distinguish between PASC and Lyme disease
Two new features were generated to confirm the distinction between CLD and LC/PASC patients after screening and to reduce classification errors. These features are part of the Lyme Index. To develop these features, we implemented an approach based on immunological importance and domain expertise. Features were generated by a programmatic method that implemented a combination of different operations (ratio, exponentiation, multiplication, sum) on the features. The approach focused on placing the important cytokines for CLD in the numerator and the important cytokines for LC/PASC in the denominator. The generated feature set was filtered to remove potential zero division and curated by domain experts to confirm biological relevance.
To determine the Lyme Index's ability to classify CLD patients, we used a dataset of 25 randomly selected CLD patients. A decision tree was trained using two features and tested on the dataset. Since there was only one class (CLD), we only calculated sensitivity, PPV (precision), and accuracy.
Predict patient outcomes based on blinded records using the best performing model
To determine the predictive ability of the best performing model upon deployment, 125 randomly selected individuals were enrolled. Individuals were processed to identify clinical assessment data and medical condition (NP, LC/PASC, or CLD) was confirmed. Individuals in the dataset were confirmed to belong to either LC/PASC or CLD medical conditions. Patients without clinical assessment data or with a diagnosis different from the model's class were excluded in order to properly calculate performance metrics. These criteria excluded only one patient profile. The resulting dataset consisted of 124 individuals, 18 CLD, and 106 LC/PASC patients defined with the same criteria as the patients in the test set, following the methods described above for LC/PASC and CLD.4,13The independent dataset consisted of groups of individuals with different ages, sexes and predominant symptoms, as summarized in Table 2.
ethics
All patients/participants provided written informed consent to participate in this study, which was approved by the IRB of the Chronic COVID Treatment Center.