Christian Magori talks about his great experience attending the Disease Ecology section of the Ecological Society of America’s 2024 Annual Meeting in the City of Angels and the fascinating things he saw and learned there.

It's been a long time since I attended the annual meeting of the Ecological Society of America in person. For the past 10 years since I became a tenured faculty member at Eastern Washington University, a regional comprehensive public university in the United States, I had to focus primarily on developing and teaching my own classes, which accounted for 80% of my workload. Attending the conference in person also comes with costs, including registration fees, airfare, accommodation, and food. It also puts a strain on my family, including losing time with my children and putting a strain on my spouse who stays home with them. Ultimately, I just didn't have the time or money to attend in person. Last summer, I tried to attend the conference virtually, but it was not a satisfying experience. However, this summer, thanks to an internal grant from my institution, I had the good fortune to attend and present at the annual meeting in person in Los Angeles.
The Ecological Society of America is the apex organization for all ecologists in the United States, and its annual meeting is the largest gathering of ecologists, usually with up to 5,000 attendees. It took place over a week from August 4 to August 9, and included workshops, invited and special sessions, contributed talks, poster presentations and networking events, social events, site visits, career development and training opportunities, and of course plenary talks. With up to 25 sessions running simultaneously from Monday afternoon to Thursday evening, I had to carefully choose which talks to attend and carefully study the schedule to plan my day. As I am a disease ecologist, I mainly attended the disease and epidemiology sessions on Wednesday and Thursday.

Hawaiian Ohia Tree (by Jana Tralut)
I made some general observations. First, the session was attended by about 100 people (the disease ecology community). The participants were a wide mix of the academic range, from undergraduates to graduate students, postdocs, and junior and senior faculty. This also reflected the academic stage of the presenters, most of the presenters were PhD students working in the labs of faculty members I still know well, but postdocs, faculty, and senior faculty also presented. In addition, some speakers worked for government agencies and non-profits. Finally, there were several undergraduates and high school students among the presenters. A great variety of approaches were presented, from pure mathematical models to statistics, field observations, molecular biology, and even social science. Topics ranged from human and animal pathogens (e.g. hantavirus, influenza, dengue, SARS-CoV-2, West Nile virus, Pd) to plant microbiomes and pathogens, such as BYDV and fungal pathogens of grasses. Ceratocystis lukuohiaIt caused rapid ohia deaths in Hawaii.The relevance of the trade-off between climate change, transmission, and virulence was one of the common themes in many of the talks I attended.
One of the biggest benefits of attending this conference for me was learning about many new developments that I can share with my students in my classes for the next few years. For example, I didn't know that there are a wide variety of databases on pathogens and their hosts and their characteristics that students can search and use everywhere, including at EWU. I had heard of the USAID PREDICT database before, but through a special session organized by the Verena Institute, I learned about the ZOVER database on zoonotic viruses, the VIRION database on vertebrate virus networks, and the PanTHERIA dataset on mammals, all of which are being used to predict and prevent the next pandemic. In the interesting discussion that followed, the presenters noted that one of the challenges of their research is the lack of longitudinal datasets on pathogen exposure and infection status over time in the same individuals (although I think veterinarians, farmers, zookeepers, and animal shelter and rescue personnel would be good sources of these observations for livestock and poultry in general). I also learned separately that a Lyme disease vaccine similar to the oral rabies vaccine is being tested to prevent infection in mice. We received great feedback on graduate student Sarah Flores' presentation about her research on seasonal prevalence of trophically transmitted parasites in invasive fish in national wildlife refuges. From how to combine different sources of information to estimate West Nile Virus risk to potential ways to control ticks in wild environments, we gained some cool and useful ideas for future research directions in the lab. Finally, I was able to reconnect with old friends and colleagues and make new connections that may lead to new opportunities and funding for myself and my students.
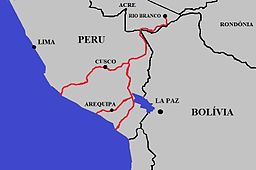
Maritime Highway Linking Peru and Brazil
It is impossible to adequately describe the 44 talks I watched and their authors in one blog post. Instead, I would like to introduce three laboratories that subjectively made a big impression on me. I was most struck by Erin Mordecai from Stanford University, who spoke about the impact of land use and land cover change on the distribution and spread of vector-borne diseases in South America. First, she shared findings linking the construction of a marine highway in Peru to increased importation and transmission of dengue fever, while correcting for many covariates, using a “difference in difference” approach to causal inference. She also linked increases in gold mining in Brazil under the Bolsonaro administration to increases in malaria cases, especially among the Yanomami, using the same methodology that she encouraged others in the audience to use. With her persuasion, I was able to look at the methodology used by social scientists and understand how it is different from, and potentially superior to, the usual multiple and partial regression methods. Second, I was surprised by the diversity of research done in Joe Hoyt’s laboratory at Virginia Tech. Here, I will focus on the study of SARS-Cov-2 in both captive wild animals in rehabilitation centers and actively captured wild animals. He and his students found the virus in a wide variety of wild animals (including deer mice, possums and raccoons) and seroprevalence increased as human presence increased, but the virus strains they detected were representative of those circulating among humans and did not suggest that new variants lurking in wildlife were reverting to humans.
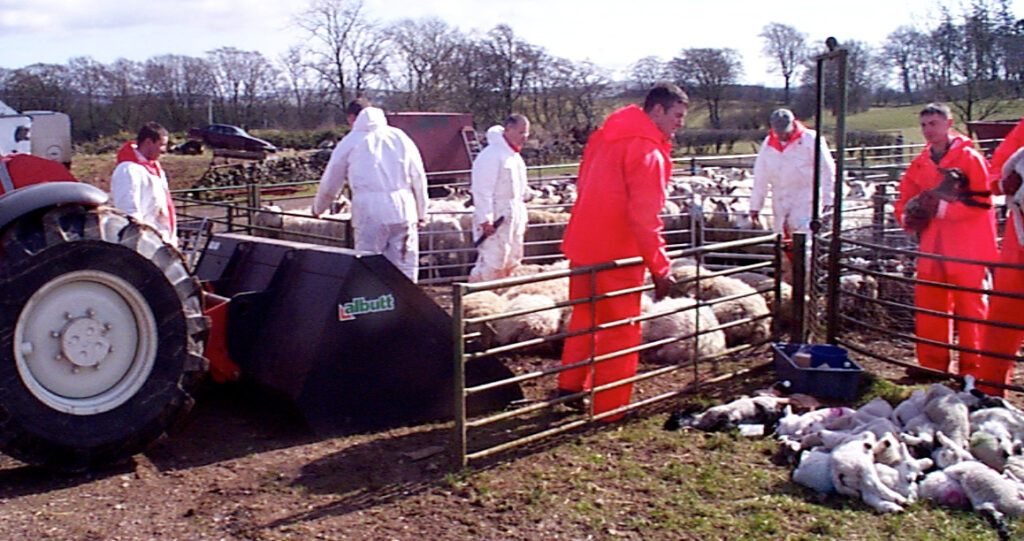

Culling of ewes and lambs in the UK in 2001 (Photo by NADIS)
Finally, I would like to introduce a talk by Ariel Greiner, a Matt Ferrari Postdoctoral Fellow at Pennsylvania State University. Greiner modeled the best surveillance methods to detect and control FMD in cattle with limited resources based on a unique dataset from the Republic of Turkey. This dataset contains information on the distance and connections between cattle farms in the country and the number of outbreaks at each farm from 2007 to 2012, and uses a time-varying network. Greiner found that focusing on a small percentage of the most connected farms (say 20%) is as good as focusing on the farms closest to previous outbreaks, or the farms most connected to previous outbreak farms, compared to random sampling. What I find great about this result is that it can be used without disease data, based only on knowing the degree of connectivity between farms. Therefore, it can be used to build surveillance systems that are 2-4 times more efficient than random sampling when there are no new disease outbreaks or when outbreaks are expected. I sincerely hope that regulatory agencies such as the USDA take note of her research and put it into practice.
Obviously, I was only able to scratch the surface of what I experienced at ESA this year, and it was also just a small part of everything it had to offer. If you're interested, you can see the full schedule and all the conference abstracts here. I hope I was able to convey how much I enjoyed it and how grateful I was to have had the opportunity to attend and present. I truly hope to return and encourage you to do the same. I look forward to seeing you all in Baltimore next year.