× close
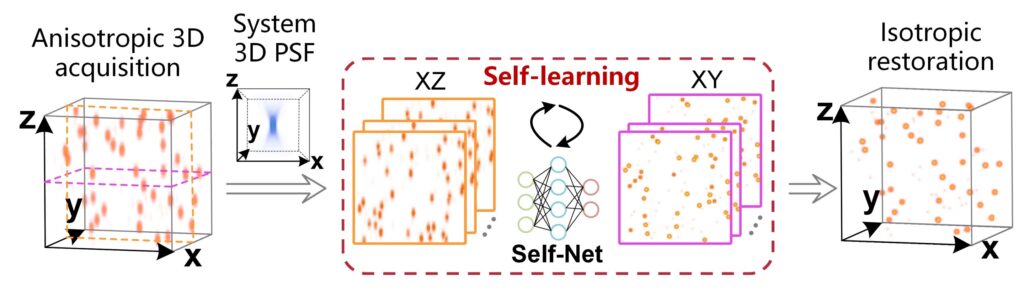
Due to the inherent anisotropy of optical PSF, HR lateral images naturally serve as a reasonable gold standard for increasing the axial resolution of raw data. Unpaired lateral and axial images obtained directly from the same anisotropic volume are used for unsupervised self-learning. Isotropic restoration is then achieved by loading the trained self-net to enhance all axial slices in the original image stack. Credits: Kefu Ning, Bolin Lu, Xiaojun Wang, Xiaoyu Zhang, Shuo Nie, Tao Jiang, Anan Li, Guoqing Fan, Xiaofeng Wang, Qingming Luo, Hui Gong, Jing Yuan
Volumetric fluorescence microscopy is an essential tool for comprehensive studies of cells and organs. Because specimens are three-dimensional (3D) in nature, an optimal imaging system should have high spatial resolution in all directions.
However, due to limitations due to their operating principles, most microscopic imaging modalities are subject to anisotropic point spread functions (PSFs). This means that axial resolution is 2-3 times worse than lateral resolution, which greatly impedes accurate visualization and dissection. Analysis of complex volumetric structures inside biological samples.
Hardware solutions to this problem are often very complex, ultimately limiting the applicability of such methods. Therefore, in current life science research, there is an urgent need to develop a simple, effective, and widely applicable 3D resolution isotropic restoration method so that 3D imaging data can be observed with consistent high resolution from different perspectives. It has become.
Unlike previous approaches, Self-Net employs unsupervised learning for realistic anisotropic degradation, thus enabling the construction of supervised training that imposes strong constraints on isotropic restoration results, resulting in high fidelity Guaranteed degree of reconstruction. This two-stage learning framework effectively alleviates the hallucination problem of previously reported methods and significantly improves the image quality.
The authors validated their method on a variety of commonly used imaging systems and biological samples. The results showed that it can not only effectively improve the 3D resolution isotropy of wide-field confocal two-photon light sheet microscopy, but also can be applied to stimulated emission depletion (STED) microscopy and super-resolution structured illumination microscopy ( SR-SIM) to achieve isotropic 3D super-resolution beyond the resolution limits of the original system.
In a paper published in Light: Science and ApplicationsA team of scientists led by Professor Hui Gong and Professor Jing Yuan from the Britton Center for Biomedical Photonics at the Wuhan National Institute of Optoelectronics, Huazhong University of Science and Technology, and their colleagues have developed a general-purpose two-stage optical sensor. A deep self-learning approach called Self-Net improves the resolution isotropy of volumetric fluorescence microscopy, provides faster training and inference speeds, and provides higher reconstruction fidelity.
The method employs a self-learning strategy to improve the axial resolution of anisotropic raw data by using high-resolution side images in the same dataset as rational targets. The unique feature of this strategy is that it takes full advantage of the 3D distribution properties of the system PSF and eliminates the need to acquire registered training data or physically model the image formation process. Based on this method, isotropic 3D imaging is easy to achieve across different microscopy platforms and is expected to facilitate the discovery of new biological insights.
The authors further demonstrated the effectiveness of Self-Net on dense neurite image data. Axonal axonal cells (AACs), one of the best characterized interneurons, contain extremely dense axonal trees, and morphological reconstructions of these cells have been shown to be useful for volumetric imaging. It has been limited by its low resolution.
With the help of Self-Net, this problem is no longer a headache. We show that the finely intermingled axonal fibers of the AAC were indistinguishable in the raw axial view but were easily resolved after self-net repair. The isotropy ratio of the 3D data increased from 0.51 to 0.94. This means that the image quality is fairly consistent even at different viewing angles. The isotropic 3D resolution provided by Self-Net allows complex 3D data in AAC to be clearly visualized and accurately analyzed. Neuron reconstruction speed has been improved nearly four times compared to previous models, and analysis efficiency has been significantly improved.
Furthermore, the high effectiveness and applicability of Self-Net allows us to achieve isotropic whole-brain imaging (DeepIsoBrain) with a voxel resolution of 0.2 × 0.2 × 0.2 μm through a time- and data-efficient pipeline.3This significantly improves the reconstruction efficiency and accuracy of complex single-neuron morphologies and could be a promising approach to facilitate morphology-based neuroscience research.
Overall, the Self-Net deep learning framework is an effective and reliable approach to achieve resolution isotropy in volume microscopy. Self-Net is expected to be useful in a variety of fields to improve existing 3D image datasets as well as new data collection.
For biological organs and tissues with unique 3D structural distributions, the isotropic 3D resolution provided by this technique effectively facilitates accurate 3D reconstruction and analysis of samples, making it useful in life sciences and other fields. It is hoped that new discoveries and new advances will be encouraged.
For more information:
Kefu Ning et al. Deep self-learning enables fast and high-fidelity isotropic resolution recovery for volumetric fluorescence microscopy. Light: Science and Applications (2023). DOI: 10.1038/s41377-023-01230-2
Magazine information:
Light: Science and Applications