patient registration
A total of 48 cases of placenta previa were enrolled, all of which were delivered at our hospital’s tertiary perinatal care center between 2008 and 2022. All patients were diagnosed with placenta previa by transvaginal ultrasound and underwent preoperative MRI upon admission. In this study, severe bleeding was defined as intraoperative blood ≥2000 ml or more. Bleeding was defined as the sum of the amount of blood lost through the gauze and suction tube.
Inclusion criteria for women was cesarean section due to diagnosis of placenta previa. An emergency caesarean section was also performed. Women who underwent both a pelvic MRI scan and a cesarean section at our institute were also included. Exclusion criteria were caesarean section in which a planned hysterectomy was performed without removal of the placenta. Cases with twins, cases with delivery at less than 22 weeks, and cases with missing patient information were also excluded (Figure 1). This study was approved by the Institutional Review Board (IRB) of Tokyo Women’s Medical University. Because this study was retrospective, the Tokyo Women’s Medical University Institutional Review Board waived the need to obtain informed consent. This study was designed and conducted in accordance with the relevant guidelines and regulations of the ethical principles for medical research involving human subjects, as described in the WMA Declaration of Helsinki.
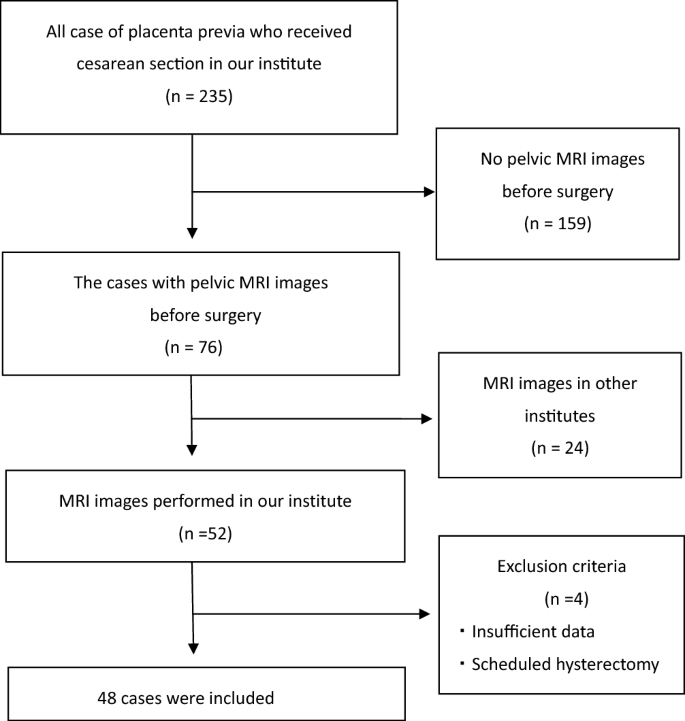
Flowchart of patient selection.
Clinical variables, MRI images, prediction targets
Two types of data were used as input data for the model: tabular data, such as patient demographics and blood tests, and imaging data, such as preoperative MRI images.
The tabular data included nine variables: (1) age, (2) number of pregnancies (gravity), (3) parity (parity), and (4) previous dynasty. The number of incisions, (5) gestational age at birth, and pre-cesarean section blood tests measure (6) birth weight, (7) hemoglobin, (8) white blood cell count, and (9) platelet count. Performing statistical analysis on each variable, he divided the patients into two groups. For statistical comparisons of two groups, Student’s t test was used to analyze significant differences in quantitative parameters. Statistical significance was set as follows. p< 0.05. We also calculated the Pearson correlation coefficient of each variable on the bleeding outcome to measure the influence of the variables.
Pelvic MRI was performed at our hospital using a 3.0-T scanner (Achieva; Philips Medical Systems, Netherlands). As previously mentioned, his MRI images from other institutions were excluded due to their impact on digital images under various conditions. Among his series of MRI images, we used sagittal MRI T2-weighted images (T2WI), which are frequently used by obstetricians in clinical practice. For each patient he used one image, and the selection criteria for one image were as follows: 1) Contains the lumen of the uterus. 2) If the uterine lumen cannot be detected, images that include the entire cervix are included. Images were selected by an obstetric expert (MA) and subsequently quality checked by another obstetric expert (KH). During this image selection process, the human experts were blinded to which images belonged to cases with intraoperative blood loss >2000 ml. After selecting a single image, a region of interest (ROI) was manually extracted using a bounding box. As her ROI for this study, we extracted a rectangular area containing the entire placenta, the intrauterine os, and the myometrium to which the placenta is attached (Figure 2). We believe that both the placenta and myometrium may have characteristics associated with bleeding. These MRI T2-weighted images (T2WI) sagittal images were divided into training and validation data in a ratio of 8:2.
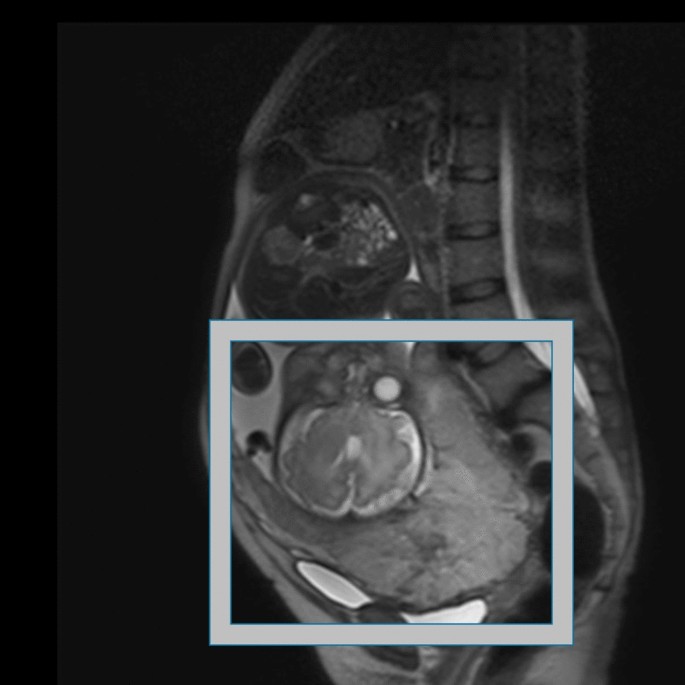
An example of an MRI image of placenta previa. In this case, the placenta covered the anterior wall of the uterus and the cervix, likely causing heavy bleeding during the caesarean section. However, the blood loss in this case was less than 2000ml. In building the model, fields indicated by white bounding boxes were used for training.
For prediction purposes, severe bleeding was defined as blood loss of 2000 mL or more at loss. The blood loss due to the suction tube and the number of gauze sheets used were calculated as the blood loss during cesarean section.
Building the model
We built a multimodal deep learning model (Model 1, shown in Figure 3) consisting of two neural networks: a convolutional neural network for MRI images and a two-layer neural network for tabular data. It takes two inputs from these neural networks and an additional neural network outputs a predicted result. For MRI images, we built a VGG16-based model as a pre-trained convolutional neural network to extract image features and output 256 vectors. For tabular data, the input was 12 variables and the output was 6 vectors through 2 hidden layers. In feature fusion, we combined and concatenated two one-dimensional vectors extracted from a convolutional neural network (CNN) model with tabular data processed by the neural network. After concatenating these two vectors, we added a three-layer neural network to generate the prediction output. For hyperparameters, we used modified linear units as the activation function, cross-entropy loss as the loss function, and stochastic gradient descent as the optimization method. The input image data was 224 × 224, and the batch size was set to 32. The CNN model was trained over 100 epochs using the early stopping method using pre-trained VGG16 weights (ImageNet dataset).
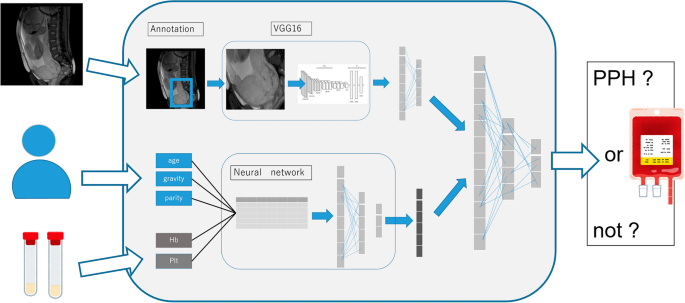
Multimodal deep learning model pipeline (Model 1).
Along with the multimodal deep learning model, we built two models using only tabular and image data (models 2 and 3 shown in Figure 4). For tabular data, we built a machine learning model based on a gradient boosting machine (XGboost). We also built his VGG16-based model for MRI images. All data were randomly split into training and validation data in an 8:2 ratio, and cross-validation was used for 1:4 cross-validation. For the simple VGG16-based model, the hyperparameters were described in a similar way to the multimodal deep learning model. For machine learning models using XGboost, the hyperparameters are: max_Depth, 4. Learning rate, 0.1; the objective was binary logistic regression and the corresponding learning objective was binary logistic regression.
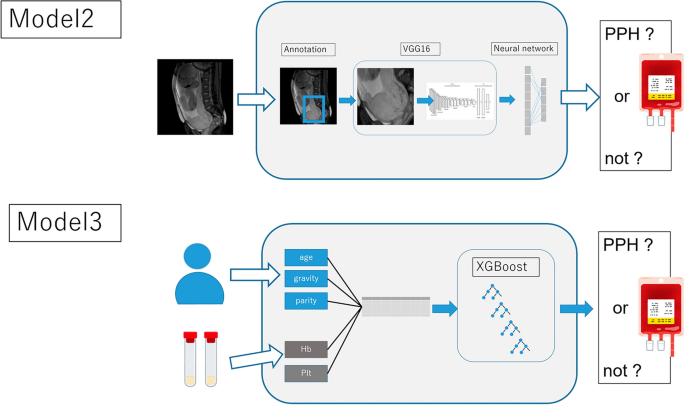
A two-model pipeline (Model 2 and Model 3) that uses only tabular and image data.
evaluation
To evaluate the variables in the tabular data, we calculated the Pearson correlation coefficient of each variable on the bleeding outcome to measure the influence of the variable. The Pearson correlation coefficient for each variable was calculated as follows.
$$ {\text{r}} = \frac{{{\text{n}}\left( {\sum {{\text{xy}}} } \right) – \left( {\sum {\text {x}} } \right)\left( {\sum {\text{y}} } \right)}}{{\left[ {{\text{n}}\sum {{\text{x}}^{2} } – \left( {\sum {\text{x}} } \right)^{2} } \right]\left[ {{\text{n}}\sum {{\text{y}}^{2} } – \left( {\sum {\text{y}} } \right)^{2} } \right]}} $$
r = Pearson coefficient, n = number of pairs of stocks, ∑xy = sum of products of pairs of stocks, ∑x = sum of x scores, ∑y = sum of y scores, ∑x2= sum of squared x scores, ∑y2= sum of squared y scores
Prediction accuracy was evaluated in terms of class accuracy and area under the curve (AUC). Class accuracy was calculated as follows.
$$ {\text{A}}\;{\text{class}}\;{\text{precision }} = \, \left( {{\text{TP}} + {\text{TN}}} \right) \, / \, \left( {{\text{TP}} + {\text{TN}} + {\text{FP}} + {\text{FN}}} \right) $$
TP, true positive; TN, true negative; FP, false positive; FN, false negative;
Point estimates for each metric were calculated by testing each model on the validation dataset. An empirical bootstrap method with 1000 iterations was used to generate 95% confidence intervals and p-values.student t-Test was used to analyze significant differences in quantitative parameters. Statistical significance was set as follows. p< 0.05. Statistical analyzes were performed using Python (version 3.7). Machine learning and deep learning were implemented using Keras (version 1.2.2) and SKlearn (version 1.0) with Python (version 3.7) as the programming language.
Finally, we tested two of our institute’s obstetric experts, allowing them to view the MRI images and tabular data together. No training was required for the human experts. Therefore, all data were used for testing and evaluated only by class accuracy.